Can dominance genetic variance be ignored in evolutionary quantitative genetic analyses of wild populations?
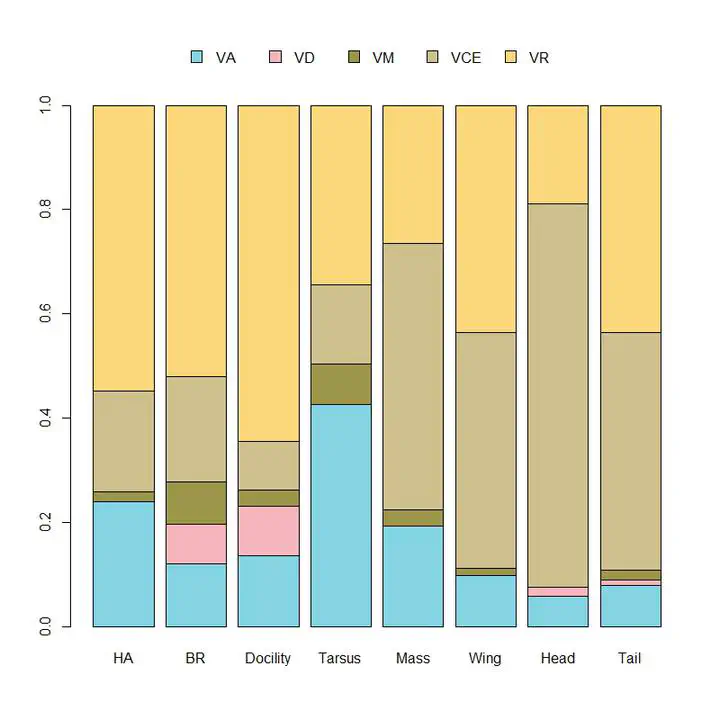 Proportions of the phenotypic variance (total column height) explained by additive genetic (VA), dominance (VD), ma- ternal (VM), common environment (VCE), and residual (VR) vari- ances calculated based on REML point estimates conditional upon the fixed effects in the model.
Proportions of the phenotypic variance (total column height) explained by additive genetic (VA), dominance (VD), ma- ternal (VM), common environment (VCE), and residual (VR) vari- ances calculated based on REML point estimates conditional upon the fixed effects in the model.Abstract
Accurately estimating genetic variance components is important for studying evolution in the wild. Empirical work on domesticated and wild outbred populations suggests that dominance genetic variance represents a substantial part of genetic variance, and theoretical work predicts that ignoring dominance can inflate estimates of additive genetic variance. Whether this issue is pervasive in natural systems is unknown, because we lack estimates of dominance variance in wild populations obtained in situ. Here, we estimate dominance and additive genetic variance, maternal variance, and other sources of nongenetic variance in eight traits measured in over 9000 wild nestlings linked through a genetically resolved pedigree. We find that dominance variance, when estimable, does not statistically differ from zero and represents a modest amount (2‐36%) of genetic variance. Simulations show that (1) inferences of all variance components for an average trait are unbiased; (2) the power to detect dominance variance is low; (3) ignoring dominance can mildly inflate additive genetic variance and heritability estimates but such inflation becomes substantial when maternal effects are also ignored. These findings hence suggest that dominance is a small source of phenotypic variance in the wild and highlight the importance of proper model construction for accurately estimating evolutionary potential.